Estimating maternal mortality rates from sibling history data
Dennis M. Feehan
2026-03-03
maternal-estimates.Rmd
library(siblingsurvival)
library(tidyverse)
#> ── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
#> ✔ dplyr 1.2.0 ✔ readr 2.2.0
#> ✔ forcats 1.0.1 ✔ stringr 1.6.0
#> ✔ ggplot2 4.0.2 ✔ tibble 3.3.1
#> ✔ lubridate 1.9.5 ✔ tidyr 1.3.2
#> ✔ purrr 1.2.1
#> ── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
#> ✖ dplyr::filter() masks stats::filter()
#> ✖ dplyr::lag() masks stats::lag()
#> ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errors
# be sure you have at least version 0.0.2.9000 of surveybootstrap
# run devtools::install_github('dfeehan/surveybootstrap')
# for the most recent version
library(surveybootstrap)
# this is helfpul for timing
library(tictoc)Overview
This vignette shows how to use the siblingsurvival
package to estimate quantities related to maternal mortality. You might
find it helpful to also read the vignette about estimating mortality
rates; much of the material here is based on that vignette. In fact,
the process for estimating maternal mortality rates is very similar to
estimating all-cause death rates, with a few modifications which we will
explain in this vignette.
It will be helpful to define a few important terms before we start our analysis:
-
ego- an ego is a survey respondent -
cell- a cell is a generic group for which we wish to produce estimates. Usually, a cell is defined by a time period, an age range, and a sex. So, for example, a cell might be women who were age 30-34 in 2015.
As an overview, we’ll start from a survey dataset, such as the one
provided by the DHS. We then use prep_dhs_sib_histories to
produce two datasets:
- one dataset has a row for each survey respondent/ego
- one dataset has a row for each sibling who is reported by a survey respondent
We’ll then calculate estimates from the sibling histories in three main steps:
- Create an esc dataset, so called because there is row for each ego X sibling X cell
- Aggregate this esc dataset up into an ec dataset, which has a row for the reports made by each ego for each cell
- Aggregate this ec dataset up into estimates for death rates, using either the individual or aggregate visibility approach (or both)
Opening up the demo datasets
We’ll start by opening up the demonstration DHS dataset.
data(model_dhs_dat)Using prep_dhs_sib_histories to get ego and sib
datasets
Now we’ll use prep_dhs_sib_histories to ready the
sibling histories for analysis. (Please see the Preparing Data vignette for more
details.) This is the step that will produce two datasets: one with a
row for each survey respondent/ego; and one with a row for each reported
sibling. Note that we include the parameter
add_maternal=TRUE to indicate that we want columns related
to maternal/pregnancy-related mortality to be added.
prepped <- prep_dhs_sib_histories(model_dhs_dat,
varmap = sibhist_varmap_dhs6,
add_maternal = TRUE,
keep_missing = FALSE)
#>
#> No information on respondent sex given; assuming all respondents are female.
#>
#> Found wwgt column; assuming we have a DHS survey and scaling weights.
#> Adding pregnancy-related/maternal death info
#> ...sib.died.accident column not found; only pregnancy-related deaths can be identified here
#> 638 out of 35082 (1.82%) reports about sibs have unknown survival status.
#> 602 out of 35082 (1.72%) reports about sibs have unknown sex.
#> Removing reported sibs missing survival status or sex.
#> ... this removes 642 out of 35082 ( 1.83 %) sibling reports.
# we'll only keep the variable we will need for this analysis
ex.ego <- prepped$ego.dat %>%
## add a 'sex' variable
mutate(sex = 'f') %>%
select(caseid,
psu,
stratum_analysis,
stratum_design,
cluster,
age.cat, sex, wwgt)
ex.sib <- prepped$sib.datLet’s take a look at the datasets we’ve produced. First, here’s a dataset that has information about survey respondents. (We’ll refer to these survey respondents as ‘ego’):
glimpse(ex.ego)
#> Rows: 8,348
#> Columns: 8
#> $ caseid <chr> " 1 1 2", " 1 3 2", " 1 4 …
#> $ psu <dbl> 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,…
#> $ stratum_analysis <dbl> 26, 26, 26, 26, 26, 26, 26, 26, 26, 26, 26, 26, 26, 2…
#> $ stratum_design <dbl> 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4,…
#> $ cluster <dbl> 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,…
#> $ age.cat <fct> "[30,35)", "[20,25)", "[40,45)", "[25,30)", "[25,30)"…
#> $ sex <chr> "f", "f", "f", "f", "f", "f", "f", "f", "f", "f", "f"…
#> $ wwgt <dbl> 1.057703, 1.057703, 1.057703, 1.057703, 1.057703, 1.0…And here’s a long-form version of the sibling history data – there’s one row for each reported sibling.
glimpse(ex.sib)
#> Rows: 34,440
#> Columns: 27
#> $ .tmpid <chr> " 1 1 2", " 1 3 2", " …
#> $ caseid <chr> " 1 1 2", " 1 3 2", " …
#> $ wwgt <dbl> 1.057703, 1.057703, 1.057703, 1.057703, 1.…
#> $ psu <dbl> 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, …
#> $ doi <dbl> 1386, 1386, 1386, 1386, 1386, 1386, 1386, …
#> $ sex <chr> "f", "f", "f", "f", "f", "f", "f", "f", "f…
#> $ sibindex <dbl> 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, …
#> $ sib.sex <chr> "m", "f", "m", "f", "m", "f", "m", "f", "m…
#> $ sib.alive <dbl> 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 0, 0, 1, …
#> $ sib.age <dbl> 42, 27, 46, 33, 40, 49, 22, 30, 49, 50, 35…
#> $ sib.dob <dbl> 876, 1056, 828, 984, 900, 792, 1116, 1020,…
#> $ sib.marital.status <dbl> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA…
#> $ sib.death.yrsago <dbl> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA…
#> $ sib.death.age <dbl> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA…
#> $ sib.death.date <dbl> -1, -1, -1, -1, -1, -1, -1, -1, -1, -1, -1…
#> $ sib.died.pregnant <dbl> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA…
#> $ sib.died.bc.pregnancy <dbl> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA…
#> $ sib.death.cause <dbl> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA…
#> $ sib.time.delivery.death <dbl> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA…
#> $ sib.place.death <dbl> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA…
#> $ sib.num.children <dbl> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA…
#> $ sib.death.year <dbl> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA…
#> $ alternum <chr> "01", "01", "01", "01", "01", "01", "01", …
#> $ sib.endobs <dbl> 1386, 1386, 1386, 1386, 1386, 1386, 1386, …
#> $ sibid <int> 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13,…
#> $ sib.preg_related.death.date <dbl> NA, -1, NA, -1, NA, -1, NA, -1, NA, NA, -1…
#> $ sib.maternal.death.date <lgl> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, NA…Laying the groundwork
First, we’ll need to add an indicator variable for each sibling’s frame population membership. In other words, we need to create a variable that has the value 1 for each sibling who was eligible to respond to the survey, and 0 otherwise. In this example data, we’ll assume that conditions similar to a typical DHS survey hold: i.e., we’ll assume the survey design was such that siblings would have been eligible to respond if they
- are alive
- are female
- are between the ages of 15 and 49
For a given survey, these criteria will differ. You’ll have to find out what the criteria for inclusion in the frame population were in order to produce estimates from sibling histories.
ex.sib <- ex.sib %>%
mutate(in.F = as.numeric((sib.alive==1) & (sib.age >= 15) & (sib.age <= 49) & (sib.sex == 'f')))Let’s look at the distribution of frame population membership
In this example dataset, a small number of the in.F
values is missing. (In a real dataset, there could well be more.) Since
we need to be able to determine whether or not each sibling is on in the
frame population, we would drop siblings missing in.F
values from the analysis.
Finally, since we will focus on pregnancy-related deaths in this vignette, we will drop all of the male sibs. We’ll also only keep female sibs up to age 49.
Specifying cells
When we produce estimated maternal death rates from sibling histories, we usually do so for different sex X age group X time period combinations. These are called cells.
Our next step is to create an object that describes the cells that we
plan to produce death rate estimates for. This means we need to specify
the time period and age groups that we’ll be using. We’ll use the helper
function cell_config to do this:
cc_pr <- cell_config(age.groups='5yr',
time.periods='7yr_beforeinterview',
start.obs='sib.dob', # date of birth
end.obs='sib.endobs', # either the date respondent was interviewed (if sib is alive) or date of death (if sib is dead)
event='sib.preg_related.death.date', # date of death (for sibs who died of maternal causes)
age.offset='sib.dob', # date of birth
time.offset='doi', # date of interview
event.name='pr_death', # pregnancy-related death
exp.scale=1/12)We’re specifying that we want
- 5-year age groups, starting from 15 and ending at 65
- to produce estimates for the 7-year window before each survey interview (so, slightly different for each respondent)
- the
dobcolumn of our sibling dataset has the time siblings start being observed (their birthdate) - the
endobscolumn of our sibling dataset has the time siblings stop being observed (the interview or, if they’re dead, the date of death) - the
sib.preg_related.death.datecolumn of our sibling dataset has the date the sibling death is considered pregnancy-related (for maternal causes, we would usesib.maternal.death.date, but this is not available for many DHS surveys) - the
doicolumn of our sibling dataset has the date the respondent who reported about the sibling was interviewed - in our survey, times are counted in months, so we set
exp.scale=1/12to indicate that we need to divide total exposures by 12 to get years
Estimating pregnancy-related death rates
Given these preparatory steps, the sibling_estimator
function will take care of estimating pregnancy-related death rates from
the sibling histories for us.
ex_ests <- sibling_estimator(sib.dat = ex.sib,
ego.id = 'caseid', # column with the respondent id
sib.id = 'sibid', # column with sibling id
# (unique for each reported sibling)
sib.frame.indicator = 'in.F', # indicator for sibling frame population membership
sib.sex = 'sib.sex', # column with sibling's sex
cell.config=cc_pr, # cell configuration we created above
weights='wwgt') # column with the respondents' sampling weights
names(ex_ests)
#> [1] "asdr.ind" "asdr.agg" "ec.dat" "esc.dat"sibling_estimator returns a list with the results. We’ll
focus on asdr.ind, which has the individual visibility
estimates, and asdr.agg, which has the aggregate visibility
estimates.
Here are the individual visibility estimates:
ex_ests$asdr.ind
#> # A tibble: 10 × 11
#> time.period sib.sex sib.age num.hat denom.hat ind.y.F n wgt.sum asdr.hat
#> <chr> <chr> <chr> <dbl> <dbl> <dbl> <int> <dbl> <dbl>
#> 1 7yr_beforei… f [15,20) 15.5 7724. 11401. 6864 6798. 0.00200
#> 2 7yr_beforei… f [20,25) 9.46 7932. 11401. 6864 6798. 0.00119
#> 3 7yr_beforei… f [25,30) 7.17 6980. 11401. 6864 6798. 0.00103
#> 4 7yr_beforei… f [30,35) 12.9 5549. 11401. 6864 6798. 0.00233
#> 5 7yr_beforei… f [35,40) 5.94 4336. 11401. 6864 6798. 0.00137
#> 6 7yr_beforei… f [40,45) 1.63 2746. 11401. 6864 6798. 0.000594
#> 7 7yr_beforei… f [45,50) 0 1752. 11401. 6864 6798. 0
#> 8 7yr_beforei… f [50,55) 0.758 987. 11401. 6864 6798. 0.000768
#> 9 7yr_beforei… f [55,60) 0 422. 11401. 6864 6798. 0
#> 10 7yr_beforei… f [60,65) 0 88.1 11401. 6864 6798. 0
#> # ℹ 2 more variables: estimator <chr>, event.name <chr>And here are the aggregate visibility estimates
ex_ests$asdr.agg
#> # A tibble: 10 × 10
#> time.period sib.sex sib.age num.hat denom.hat n wgt.sum asdr.hat
#> <chr> <chr> <chr> <dbl> <dbl> <int> <dbl> <dbl>
#> 1 7yr_beforeint f [15,20) 29.2 14696. 6864 6798. 0.00199
#> 2 7yr_beforeint f [20,25) 24.1 15949. 6864 6798. 0.00151
#> 3 7yr_beforeint f [25,30) 18.2 14467. 6864 6798. 0.00126
#> 4 7yr_beforeint f [30,35) 22.8 11597. 6864 6798. 0.00197
#> 5 7yr_beforeint f [35,40) 12.8 8852. 6864 6798. 0.00144
#> 6 7yr_beforeint f [40,45) 2.61 5321. 6864 6798. 0.000491
#> 7 7yr_beforeint f [45,50) 0 3066. 6864 6798. 0
#> 8 7yr_beforeint f [50,55) 0.758 1573. 6864 6798. 0.000482
#> 9 7yr_beforeint f [55,60) 0 609. 6864 6798. 0
#> 10 7yr_beforeint f [60,65) 0 111. 6864 6798. 0
#> # ℹ 2 more variables: estimator <chr>, event.name <chr>Plotting the results
We’ll make some plots showing the results. We’ll only show results up to age 50, which is typically how pregnancy-related mortality is analyzed.
ggplot(ex_ests$asdr.ind %>% filter(! sib.age %in% c("[50,55)", "[55,60)", "[60,65)"))) +
geom_line(aes(x=sib.age, y=1000*asdr.hat, group=sib.sex)) +
theme_minimal() +
scale_y_log10() +
ylab("1,000 X PRMR") +
ggtitle('individual visibility estimator\npregnancy-related mortality, 7yr before survey')
#> Warning in scale_y_log10(): log-10 transformation introduced
#> infinite values.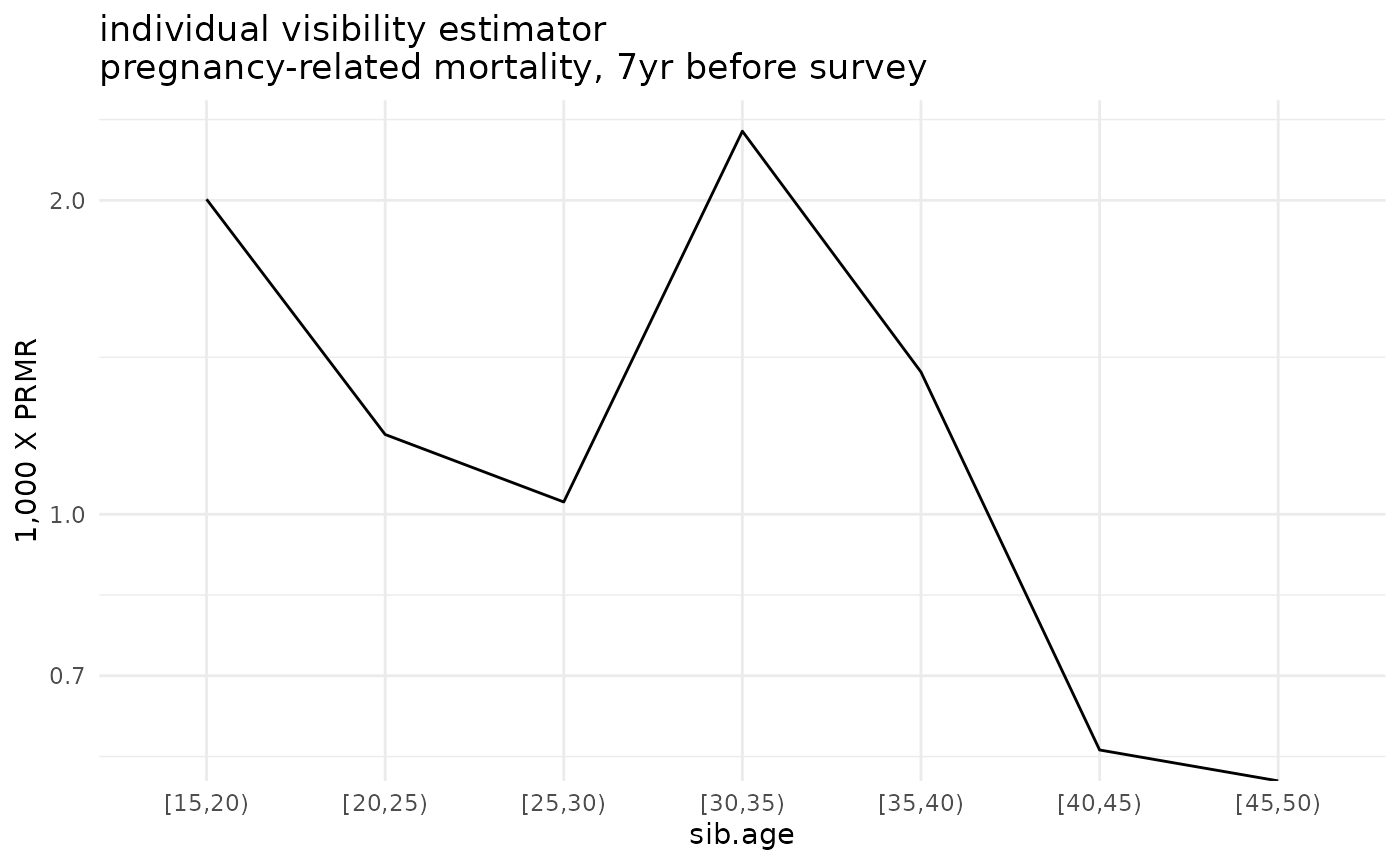
ggplot(ex_ests$asdr.agg %>% filter(! sib.age %in% c("[50,55)", "[55,60)", "[60,65)"))) +
geom_line(aes(x=sib.age, y=1000*asdr.hat, group=sib.sex)) +
theme_minimal() +
scale_y_log10() +
ylab("1,000 X PRMR") +
ggtitle('aggregate visibility estimator\npregnancy-related mortality, 7yr before survey')
#> Warning in scale_y_log10(): log-10 transformation introduced
#> infinite values.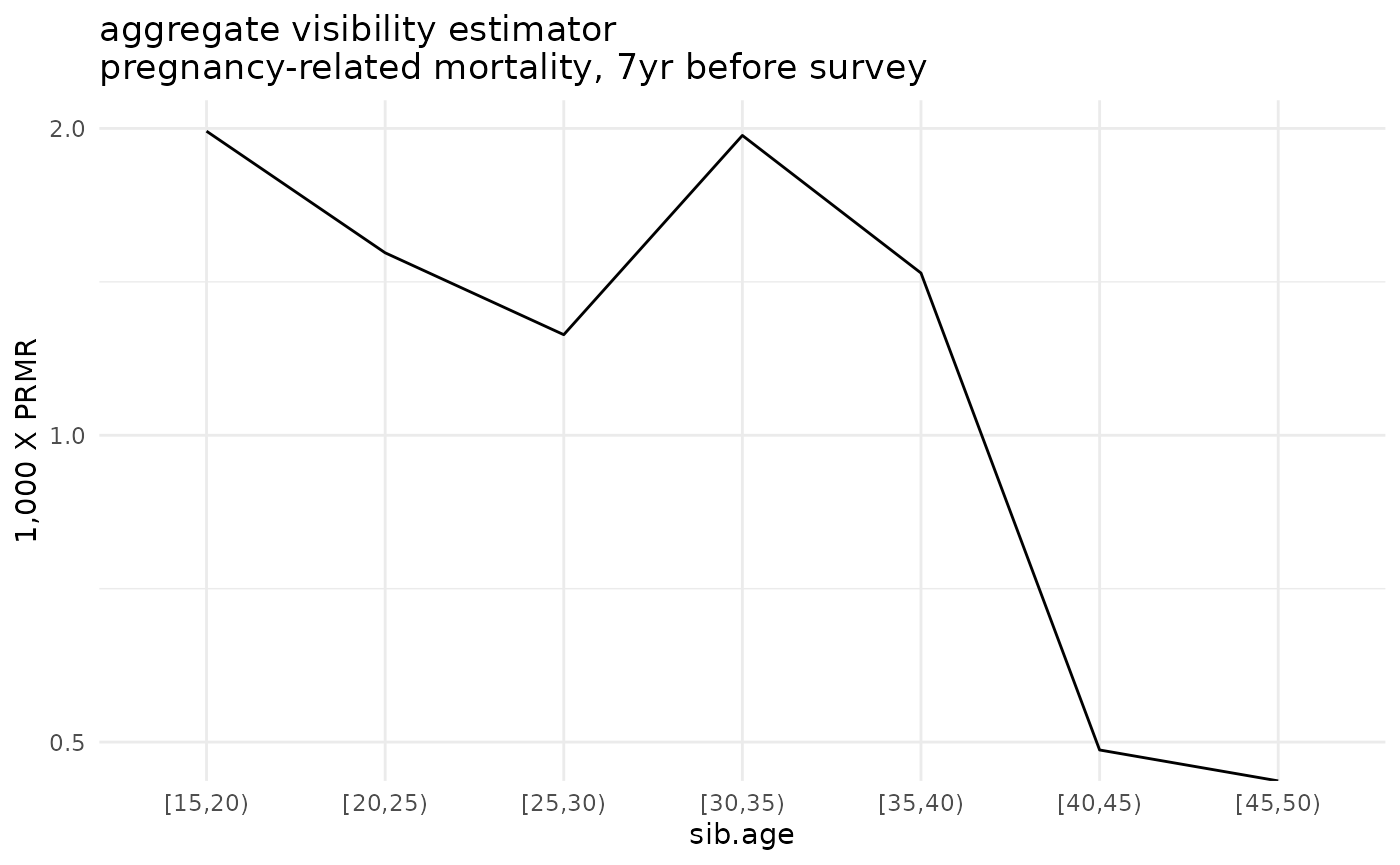
compare <- bind_rows(ex_ests$asdr.ind, ex_ests$asdr.agg)
ggplot(compare %>% filter(! sib.age %in% c("[50,55)", "[55,60)", "[60,65)"))) +
geom_line(aes(x=sib.age, y=1000*asdr.hat, linetype=estimator, group=interaction(estimator, sib.sex))) +
theme_minimal() +
#facet_grid(sex ~ .) +
facet_grid(. ~ sib.sex) +
scale_y_log10() +
ylab("1,000 X PRMR") +
ggtitle('both estimators\npregnancy-related mortality, 7yr before survey')
#> Warning in scale_y_log10(): log-10 transformation introduced
#> infinite values.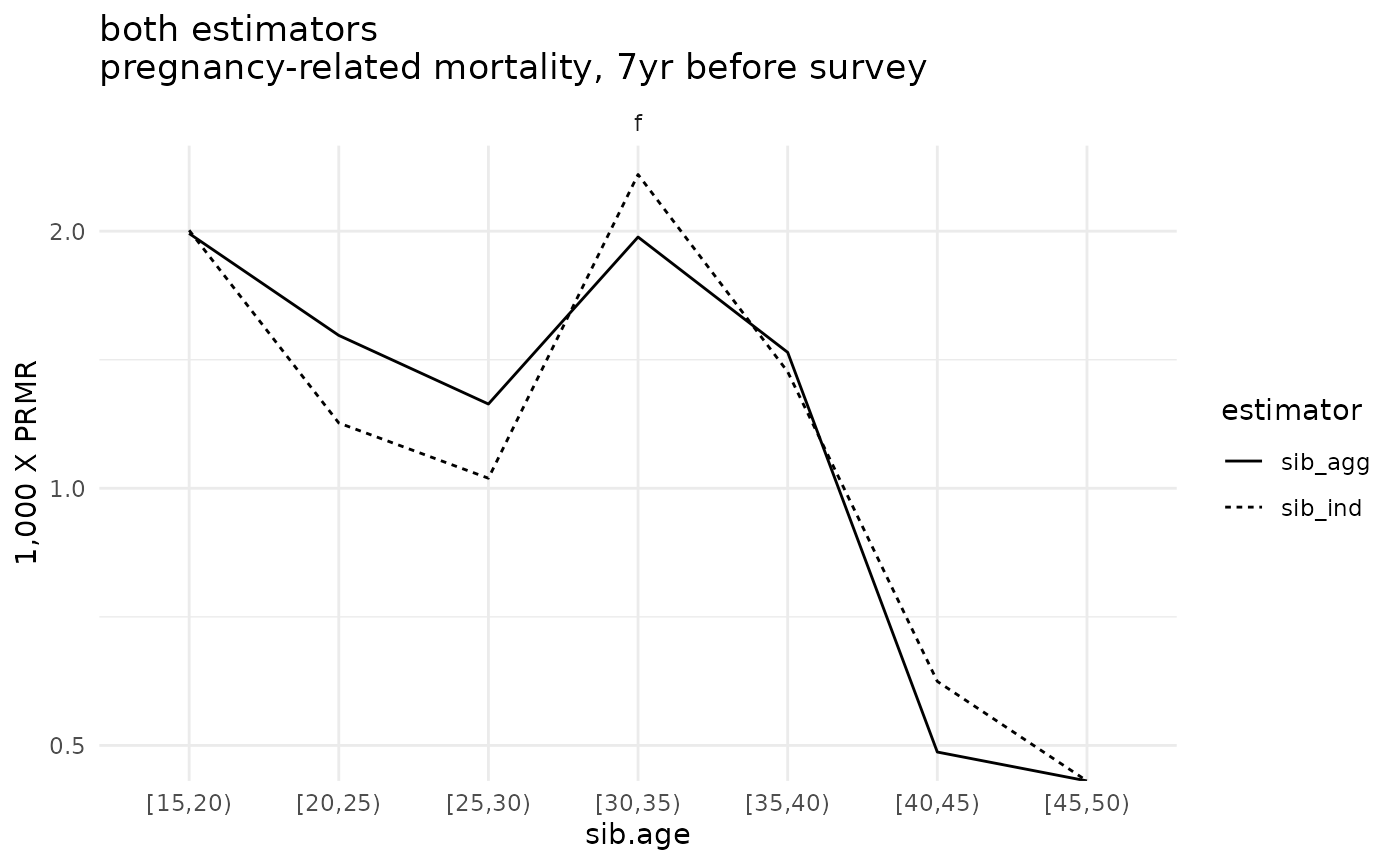
Aggregating estimates across ages
Problem: ex.sib doesn’t have column age.cat
mmrate <- aggregate_maternal_estimates(ex_ests, ex.ego, ex.sib)
#> Joining with `by = join_by(.ego.id, .weight, sex)`
#> Joining with `by = join_by(time.period, sib.sex, sib.age, event.name)`
mmrate
#> # A tibble: 1 × 7
#> ind.est agg.est adj.factor adj.factor.allage adj.factor.meanagespec
#> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 0.00138 0.00143 0.430 0.577 0.573
#> # ℹ 2 more variables: ratio.agg.ind <dbl>, ratio.ind.agg <dbl>Variance estimates
In practice, we want point estimates and estimated sampling
uncertainty for the death rates. We’ll use the rescaled bootstrap to
estimate sampling uncertainty. This can be done with the
surveybootstrap package.
Before we use the rescaled bootstrap, we need to know a little bit about the sampling design of the survey we’re working with. In this example dataset, we have a stratified, multistage design. So we’ll need to tell the bootstrap function about the strata and the primary sampling units. In this example data, these are indicated by the ‘stratum’ and ‘psu’ columns of the dataset.
(NOTE: this takes about a minute or so on a 2018 MBP for 1000 resamples.)
set.seed(101010)
tic('running bootstrap')
## this will take a little while -- for 1000 reps, it takes about 10 minutes on a 2018 Macbook Pro
#num.boot.reps <- 1000
# reduce number of reps to help vignette build faster
num.boot.reps <- 100
# The Guide to DHS Statistics DHS-8
# suggests using what we call the `stratum_design` variable
# (v023) in calculating sampling-based standard errors
bootweights <- surveybootstrap::rescaled.bootstrap.weights(survey.design = ~ psu + strata(stratum_design),
# a high number is good here, though that will obviously
# make everything take longer
num.reps=num.boot.reps,
idvar='caseid', # column with the respondent ids
weights='wwgt', # column with the sampling weight
survey.data=ex.ego # note that we pass in the respondent data, NOT the sibling data
)
#> Warning: Unquoting language objects with `!!!` is deprecated as of rlang 0.4.0. Please
#> use `!!` instead.
#>
#> # Bad: dplyr::select(data, !!!enquo(x))
#>
#> # Good: dplyr::select(data, !!enquo(x)) # Unquote single quosure
#> dplyr::select(data, !!!enquos(x)) # Splice list of quosures
#> This warning is displayed once every 8 hours.
toc()
#> running bootstrap: 0.784 sec elapsedThe result, bootweights, is a dataframe that has a row
for each survey respondent, a column for the survey respondent’s ID, and
then num.boot.reps columns containing survey weights that
result from the bootstrap procedure. The basic idea is to calculate
estimated death rates using each one of the num.boot.reps
sets of weights. The variation across the estimates is then an estimator
for the sampling variation.
To make this easier, the sibling_estimator function can
take a dataset with bootstrap resampled weights; it will then calculate
and summarize the estimates for you.
(NOTE: this is slow; it takes about 35 minutes or so on a 2018 MBP
when bootweights has 1000 resamples.)
# to save time, we'll only use a subset of the bootstrap replicates
short.bootweights <- bootweights %>% select(1:11)
#est.bootweights <- bootweights
est.bootweights <- short.bootweights
tic('calculating estimates with bootstrap')
ex_boot_ests <- sibling_estimator(sib.dat = ex.sib,
ego.id = 'caseid',
sib.id = 'sibid',
sib.frame.indicator = 'in.F',
sib.sex = 'sib.sex',
cell.config=cc_pr,
boot.weights=est.bootweights, # to get sampling uncertainty, we pass boot.weights into sibling_estimator
return.boot=TRUE, # when TRUE, return all of the resampled estimates (not just summaries)
weights='wwgt')
toc()
#> calculating estimates with bootstrap: 3.91 sec elapsedFinally, let’s plot the estimated death rates along with their sampling uncertainty:
ggplot(ex_boot_ests$asdr.ind %>% filter(! sib.age %in% c("[50,55)", "[55,60)", "[60,65)"))) +
geom_ribbon(aes(x=sib.age, ymin=1000*asdr.hat.ci.low, ymax=1000*asdr.hat.ci.high, group=sib.sex), alpha=.2) +
geom_line(aes(x=sib.age, y=1000*asdr.hat, group=sib.sex)) +
theme_minimal() +
scale_y_log10() +
ylab("1,000 X PRMR") +
ggtitle('individual visibility estimator\npregnancy-related mortality, 7yr before survey')
#> Warning in scale_y_log10(): log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.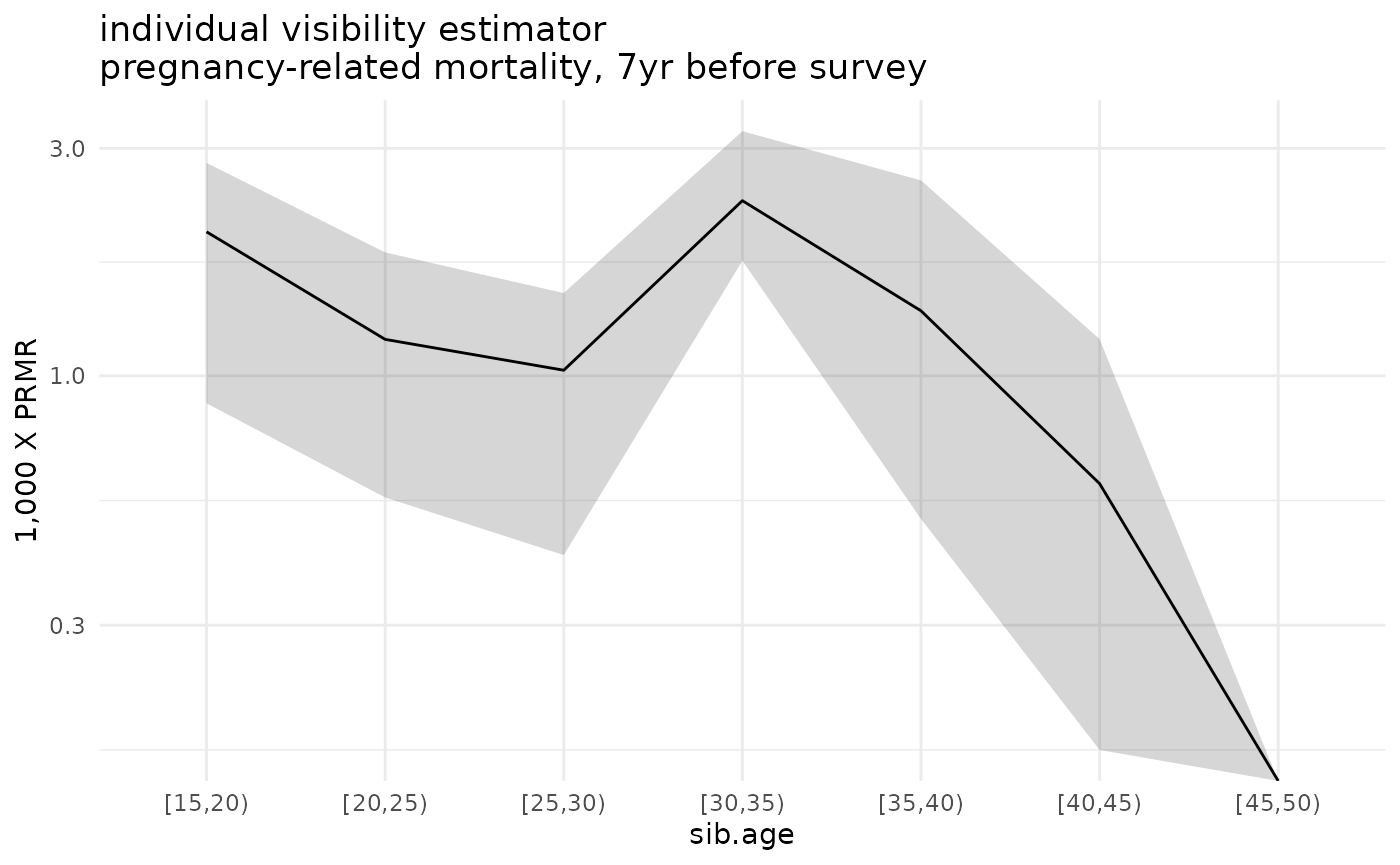
ggplot(ex_boot_ests$asdr.agg %>% filter(! sib.age %in% c("[50,55)", "[55,60)", "[60,65)"))) +
geom_ribbon(aes(x=sib.age, ymin=1000*asdr.hat.ci.low, ymax=1000*asdr.hat.ci.high, group=sib.sex), alpha=.2) +
geom_line(aes(x=sib.age, y=1000*asdr.hat, group=sib.sex)) +
theme_minimal() +
scale_y_log10() +
ylab("1,000 X PRMR") +
ggtitle('aggregate visibility estimator\npregnancy-related mortality, 7yr before survey')
#> Warning in scale_y_log10(): log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.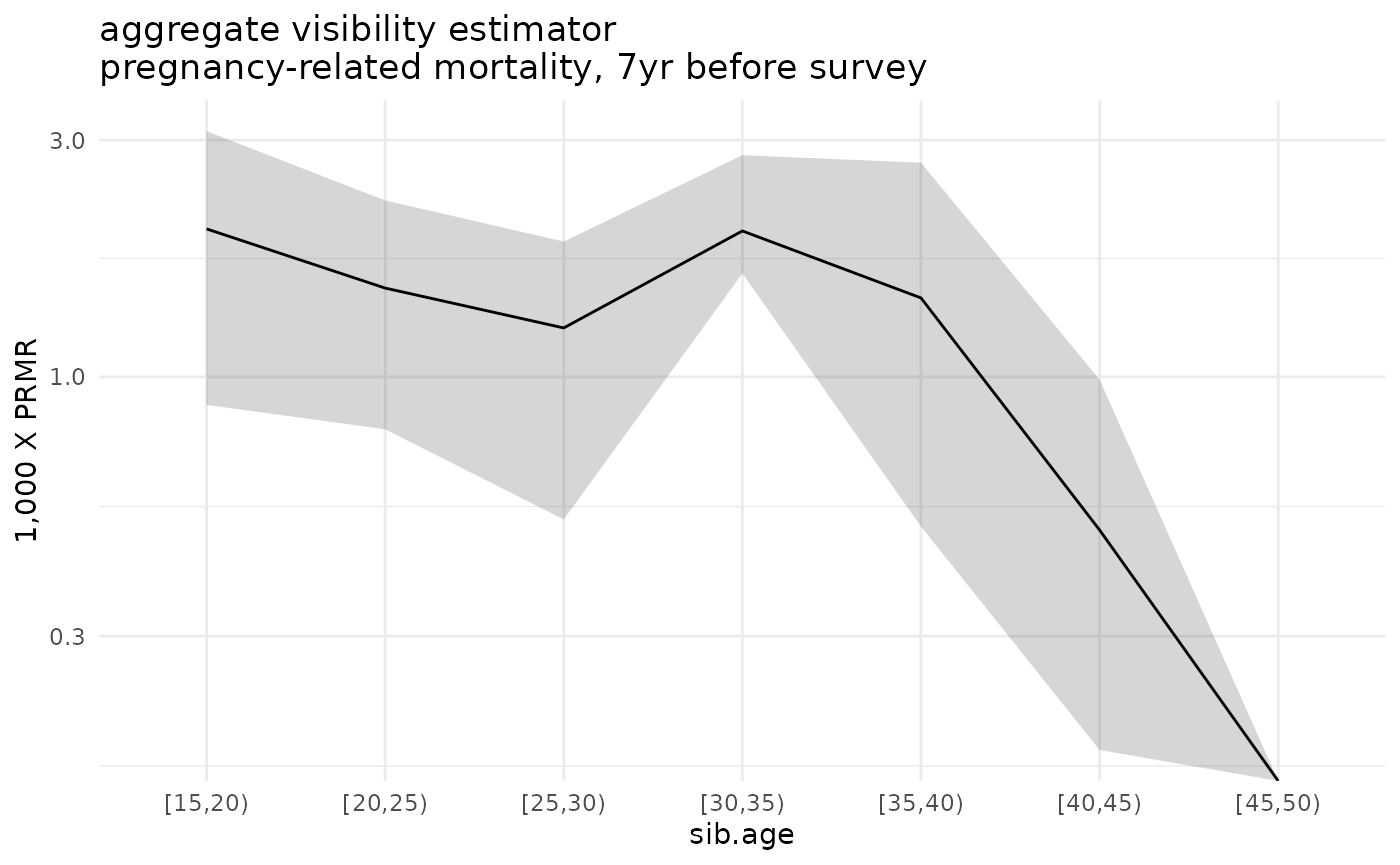
compare_boot <- bind_rows(ex_boot_ests$asdr.ind, ex_boot_ests$asdr.agg)
ggplot(compare_boot %>% filter(! sib.age %in% c("[50,55)", "[55,60)", "[60,65)"))) +
geom_ribbon(aes(x=sib.age, ymin=1000*asdr.hat.ci.low, ymax=1000*asdr.hat.ci.high, fill=estimator, group=estimator), alpha=.2) +
geom_line(aes(x=sib.age, y=1000*asdr.hat, color=estimator, group=interaction(estimator))) +
theme_minimal() +
scale_y_log10() +
ylab("1,000 X PRMR") +
ggtitle('both estimators\npregnancy-related mortality, 7yr before survey')
#> Warning in scale_y_log10(): log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.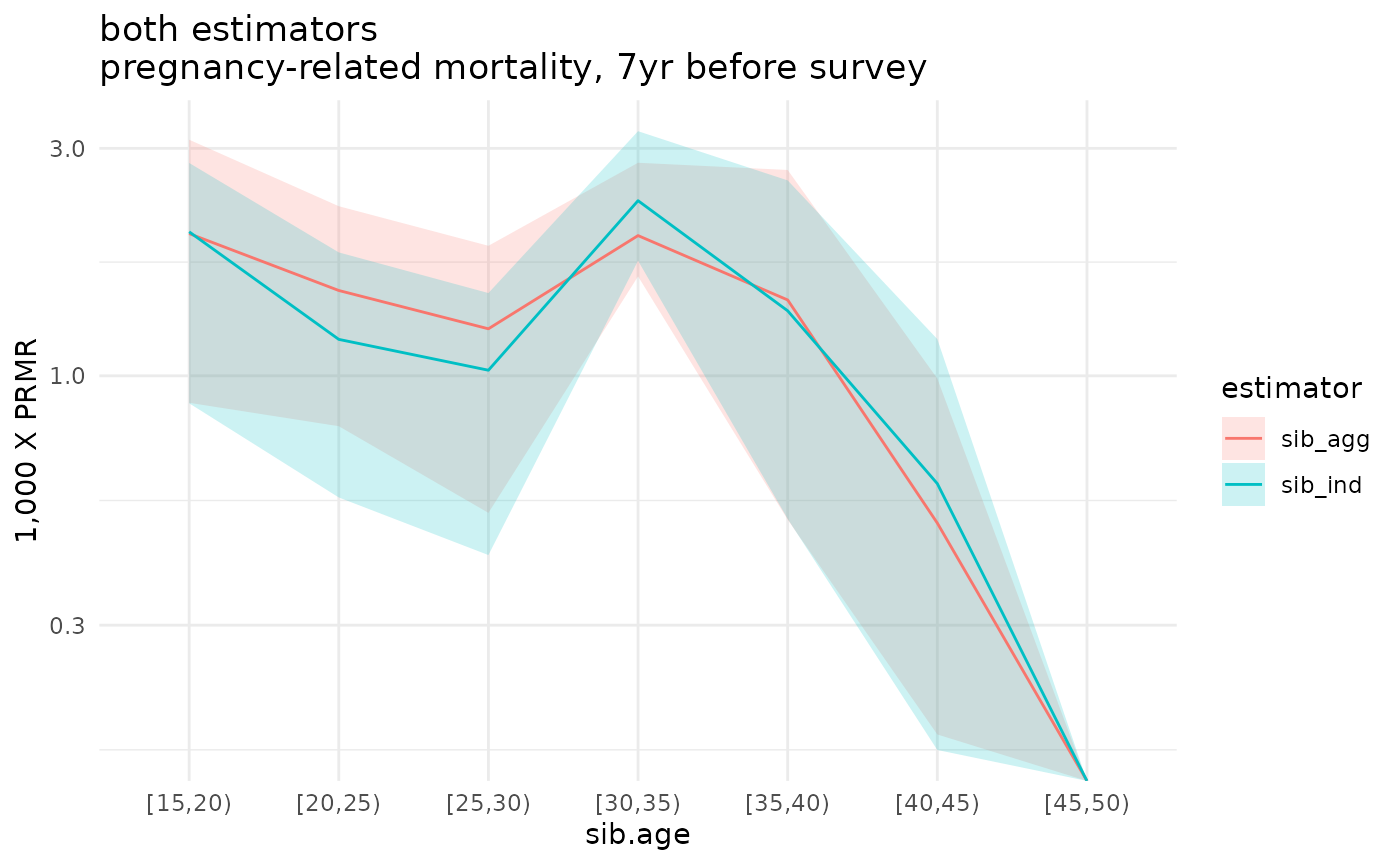
Finally, we can calculate the aggregated MMRate for pregnancy-related mortality using the same bootstrap resamples.
mmrate_boot <- aggregate_maternal_estimates(ex_boot_ests, ex.ego, ex.sib)
#> Joining with `by = join_by(.ego.id, .weight, sex)`
#> Joining with `by = join_by(time.period, sib.sex, sib.age, event.name)`
#> Joining with `by = join_by(time.period, sib.sex, sib.age, boot_idx,
#> event.name)`
names(mmrate_boot)
#> [1] "point" "boot_summ" "boot"