The rescaled bootstrap
Dennis M. Feehan
2026-03-09
Source:vignettes/rescaled_bootstrap.Rmd
rescaled_bootstrap.RmdIntroduction
This vignette demonstrates the basic functionality of the
surveybootstrap package. The goal is to illustrate how to
use the rescaled bootstrap (Rao and Wu 1988; Rust and
Rao 1996) to take bootstrap resamples from a survey dataset when the
survey sample has a complex design.
The rescaled bootstrap handles complex designs by
resampling at the level of the primary sampling unit (PSU) and adjusting
the sampling weights of every observation in each bootstrap replicate.
Specifically, for each bootstrap resample, we draw
PSUs with replacement from the
PSUs in stratum
,
and rescales the observation weights to reflect how many times each PSU
was selected.
This package implements the rescaled bootstrap in C++ (via Rcpp) for
speed.
Use rescaled.bootstrap.sample /
bootstrap.estimates when your survey has a cluster or
stratified design. For a simple random sample without clusters or
strata, use srs.bootstrap.sample instead.
The MU284 example data
The MU284 dataset (Sarndal, Swensson, and Wretman 2003,
pg. 652) describes 284 Swedish municipalities. Relevant columns
include:
| Column | Description |
|---|---|
LABEL |
Municipality ID |
S82 |
Total seats in municipal council |
RMT85 |
Municipal tax revenue, 1985 (in millions of Kronor) |
P85 |
Population, 1985 (in thousands) |
CL |
Cluster indicator (a set of neighboring municipalities) |
MU284.complex.surveys is a list of 10 sample surveys
drawn from this population using the two-stage design from Example 4.3.2
of Sarndal et al.:
- Stage I — simple random sample without replacement of PSUs (clusters) from total PSUs
- Stage II — simple random sample without replacement of municipalities from each selected PSU
This gives
sampled municipalities per survey. Each row has a
sample_weight reflecting the inverse of its selection
probability.
library(surveybootstrap)
survey <- MU284.complex.surveys[[1]]
head(survey[, c("LABEL", "CL", "S82", "RMT85", "P85", "sample_weight")])
#> LABEL CL S82 RMT85 P85 sample_weight
#> 1 96 17 41 81 13 26.66667
#> 2 99 17 49 129 20 26.66667
#> 3 101 17 61 299 49 26.66667
#> 4 16 4 101 6263 653 16.66667
#> 5 18 4 61 532 59 16.66667
#> 6 19 4 41 250 27 16.66667There are two ways to use the package:
- Path 1: specifying a function to produce and estimate from data with bootstrap weights
- Path 2: getting a matrix of bootstrap weights
Path 1: bootstrap.estimates()
bootstrap.estimates() is the highest-level interface.
You supply an estimator function and a bootstrap method; it returns
bootstrap replicate estimates that you can use to compute standard
errors and confidence intervals.
Step 1: define an estimator
Your estimator must accept two arguments — survey.data
(the resampled dataset) and weights (a vector of rescaled
sampling weights) — and return a named data frame with one row.
my_estimator <- function(survey.data, weights) {
data.frame(
total_S82 = sum(survey.data$S82 * weights),
ratio_RMT85_P85 = sum(survey.data$RMT85 * weights) /
sum(survey.data$P85 * weights)
)
}Here total_S82 is the Horvitz-Thompson estimator of the
population total of S82 (total council seats), and
ratio_RMT85_P85 is a ratio estimator (tax revenue per
capita).
Step 2: Run the bootstrap
set.seed(12345)
boot_res <- bootstrap.estimates(
survey.data = survey,
survey.design = ~ CL, # CL identifies the PSU (cluster)
num.reps = 500,
estimator.fn = my_estimator, # we just defined this above
weights = "sample_weight",
bootstrap.fn = "rescaled.bootstrap.sample"
)The survey.design formula uses the column
CL as the PSU identifier. There are no explicit strata
here, so the entire sample is treated as a single stratum. See Describing your survey design
below for formulas with strata.
boot_res is a list of 500 one-row data frames, one per
bootstrap replicate.
Step 3: Collect and summarize
boot_df <- do.call("rbind", boot_res)
# Bootstrap standard errors
apply(boot_df, 2, sd)
#> total_S82 ratio_RMT85_P85
#> 1638.2395064 0.8185707
# Bootstrap means (should be close to the original point estimates)
apply(boot_df, 2, mean)
#> total_S82 ratio_RMT85_P85
#> 15290.525000 8.367827
# Original point estimates for comparison
my_estimator(survey, survey$sample_weight)
#> total_S82 ratio_RMT85_P85
#> 1 15400 8.765395You can also pass summary.fn to
bootstrap.estimates() to get a summary computed in a single
pass without keeping all replicates in memory:
boot_summary <- bootstrap.estimates(
survey.data = survey,
survey.design = ~ CL,
num.reps = 500,
estimator.fn = my_estimator,
weights = "sample_weight",
bootstrap.fn = "rescaled.bootstrap.sample",
summary.fn = MU284.estimator.summary.fn, # example summary function
verbose = FALSE
)Visualising the bootstrap distribution
hist(boot_df$total_S82,
main = "Bootstrap distribution: total council seats",
xlab = "Estimated total (S82)",
col = "steelblue",
border = "white")
abline(v = my_estimator(survey, survey$sample_weight)$total_S82,
col = "firebrick", lwd = 2, lty = 2)
legend("topright", legend = "Point estimate", col = "firebrick",
lwd = 2, lty = 2, bty = "n")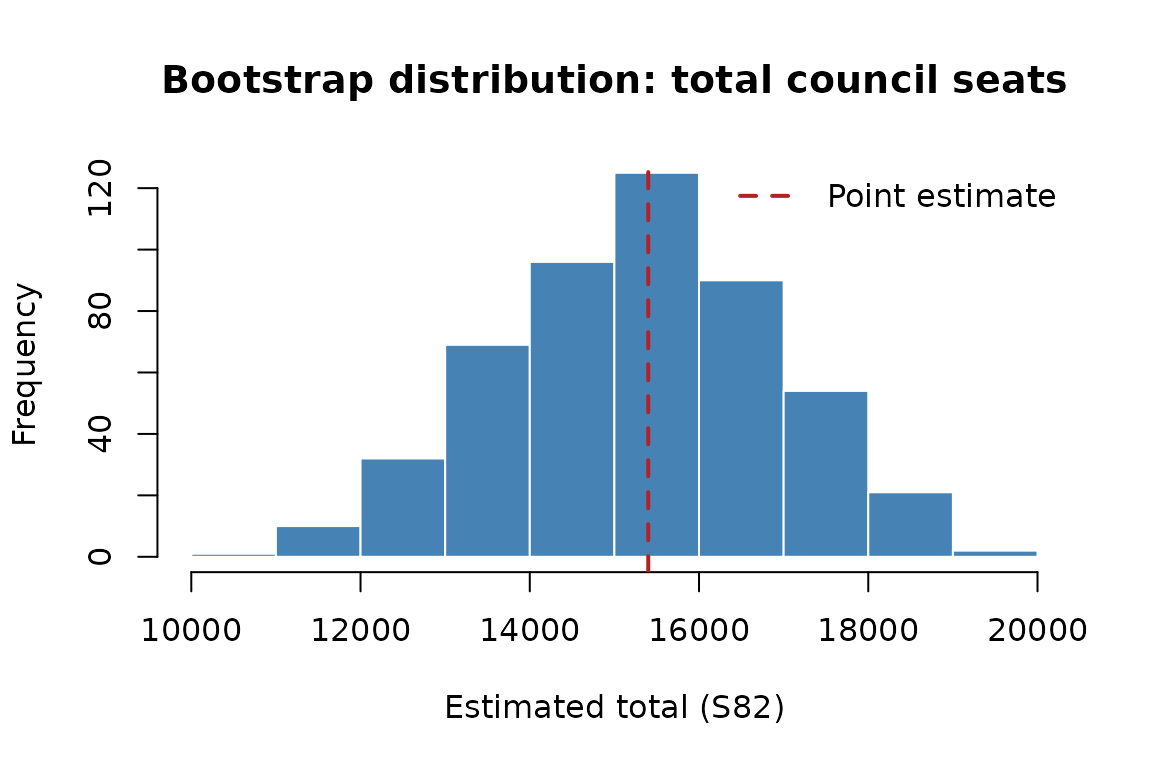
Path 2: get.rescaled.bootstrap.weights()
Sometimes it is more convenient to obtain a matrix of bootstrap weights directly and then apply your estimator(s) yourself. This is useful when:
- you want to compute many different estimators from the same set of replicates
- you need to pass bootstrap weights to external software
- you are implementing jackknife-after-bootstrap (JAB) variance estimation
Obtain bootstrap weights
boot_wts <- get.rescaled.bootstrap.weights(
survey.data = survey,
survey.design = ~ CL,
idvar = "LABEL",
weights = "sample_weight",
num.reps = 500
)The result is a list with two elements:
-
orig_weights— a data frame with one row per respondent and aweightcolumn containing the original sampling weight -
boot_weights— a data frame with one row per respondent and columnsLABEL,boot_weight_1, …,boot_weight_500
names(boot_wts)
#> [1] "orig_weights" "boot_weights"
# First few rows and columns of boot_weights
boot_wts$boot_weights[1:5, 1:4]
#> LABEL boot_weight_1 boot_weight_2 boot_weight_3
#> 1 96 0 66.66667 33.33333
#> 2 99 0 66.66667 33.33333
#> 3 101 0 66.66667 33.33333
#> 4 16 0 20.83333 20.83333
#> 5 18 0 20.83333 20.83333Compute estimates from bootstrap weights
Join the survey data to the weight matrix, then apply your estimator for each replicate:
# Merge survey data with bootstrap weights on the respondent id
bw <- merge(survey[, c("LABEL", "S82", "RMT85", "P85")],
boot_wts$boot_weights,
by = "LABEL")
# Compute the total_S82 estimator for each bootstrap replicate
boot_totals <- sapply(paste0("boot_weight_", 1:500), function(col) {
sum(bw$S82 * bw[[col]])
})
# Bootstrap standard error
sd(boot_totals)
#> [1] 1607.395Scaling factors
Pass include_scaling_factors = TRUE to also get the
per-replicate weight scaling factors. Each bootstrap weight is the
product of the original weight and the corresponding scaling factor:
boot_wts_sf <- get.rescaled.bootstrap.weights(
survey.data = survey,
survey.design = ~ CL,
idvar = "LABEL",
weights = "sample_weight",
num.reps = 500,
include_scaling_factors = TRUE
)
# boot_wts_sf$weight_scaling_factor has columns LABEL, boot_rep_1, ..., boot_rep_500Describing your survey design
The survey.design argument is a formula that tells
surveybootstrap which columns identify PSUs and strata.
| Formula | Meaning |
|---|---|
~ CL |
PSU is CL; no strata (one global stratum) |
~ psu + strata(region) |
PSU is psu; strata defined by region
|
~ psu1 + psu2 + strata(s1 + s2) |
PSU uniquely identified by (psu1, psu2); strata by
(s1, s2)
|
~ 1 |
Simple random sample — use srs.bootstrap.sample
instead |
For a simple random sample, use
bootstrap.fn = "srs.bootstrap.sample" and
survey.design = ~ 1:
srs_survey <- MU284.surveys[[1]]
boot_srs <- bootstrap.estimates(
survey.data = srs_survey,
survey.design = ~ 1,
num.reps = 500,
estimator.fn = my_estimator,
weights = "sample_weight",
bootstrap.fn = "srs.bootstrap.sample"
)References
Rao, J.N.K. and Wu, C.F.J. (1988). Resampling inference with complex survey data. Journal of the American Statistical Association, 83(401), 231–241. https://doi.org/10.1080/01621459.1988.10478591
Rao, J. N. K., C. F. J. Wu, and Kim Yue. “Some recent work on resampling methods for complex surveys.” Survey Methodology 18.2 (1992): 209-217.
Rust, K.F. and Rao, J.N.K. (1996). Variance estimation for complex surveys using replication techniques. Statistical Methods in Medical Research, 5(3), 283–310. https://doi.org/10.1177/096228029600500305
Sarndal, C.-E., Swensson, B. and Wretman, J. (2003). Model Assisted Survey Sampling. Springer. ISBN: 0387406204.